- Osirix lite segmentation manual#
- Osirix lite segmentation full#
- Osirix lite segmentation software#
- Osirix lite segmentation series#
- Osirix lite segmentation free#
Osirix lite segmentation software#
"OsiriX HD" is a DICOM software for iOS: DICOM is the digital standard for storing and transferring medical images. Mobie Awards 2009: Winner - Best Medical App Aunt Minnie's 2013 - "Best Radiology Mobile App”
Osirix lite segmentation full#
and the FCN model introduced by Tran, we designed a novel \(19-\) layer FCN for an automated pixelwise image segmentation in CHD subjects.OsiriX HD is a full DICOM image viewer for iOS (DICOM Files & DICOM Network protocol support). Inspired by the “skip” architecture used by Long et al. While these FCNs yield good segmentation accuracy for healthy adult CMR images, they show poor performance on CHD subjects. Examples of FCNs used for segmenting healthy adult CMR images include.
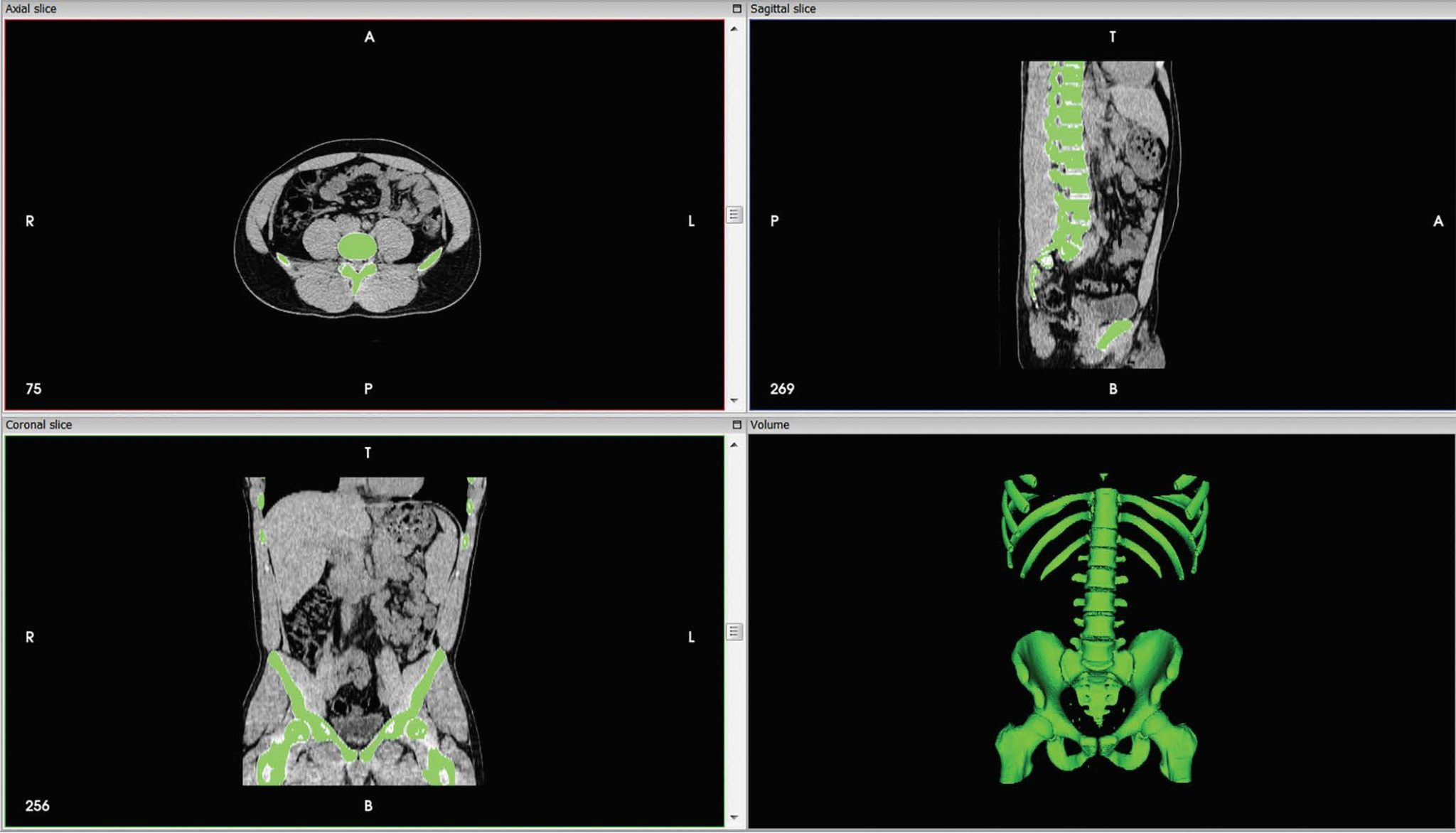
These networks are mostly useful when the data is either an image or a map such that the proximity among pixels represents how associated they are.
Osirix lite segmentation series#
Convolutional networks are a family of artificial neural networks that are comprised of a series of convolutional and pooling layers in which the data features are learned in various levels of abstraction. Image segmentation using fully convolutional networksĪ fully convolutional network (FCN), in comparison with a U-net and cvi42, was used for automated pixelwise image segmentation. For training data, twenty-six patients ( \(10\) TOFs, \(4\) DORVs, \(4\) TGAs, \(4\) CAAs and \(4\) cardiomyopathy patients) were randomly selected whereas the remaining \(38\) patients were used as test data. The entire process was performed using two different down-sampling methods: (1) nearest-neighbor down-sampling and (2) bi-cubical down-sampling.
To reduce the dimensionality, each cropped image was subsequently resized to \(128\times 128\) using the imresize function in the open-source Python library SciPy. Subsequently, all images were examined to ensure that both the heart and segmentation mask are present.
The original dataset was first preprocessed by center-cropping each image to the size of \(445\times 445\) to remove patients’ identifiers. The original image size was \(512\times 512\) pixels. The ventricular cavity in the basal slice was identified by evaluating wall thickening and cavity shrinking in systole.
Osirix lite segmentation manual#
Manual annotations were performed according to SCMR guidelines with cvi42 software (Circle Cardiovascular Imaging, Calgary, Alberta, Canada) without the use of automated segmentation tools. Ventricular volumes and ejection fraction were then computed from these contours. Endocardial contours were drawn on end-diastolic and end-systolic images.
Manual image segmentation was performed by a board-certified pediatric cardiologist sub-specialized in CMR with experience consistent with Society for Cardiovascular Magnetic Resonance (SCMR) Level 3 certification.
Osirix lite segmentation free#
Images were obtained with the patients free breathing \(3\) signal averages were obtained to compensate for respiratory motion. Resultsįor congenital CMR dataset, our FCN model yields an average Dice metric of \(91.0\mathrm\), and flip angle of \(60\) degrees. Dice metric, Jaccard index and Hausdorff distance as well as clinically-relevant volumetric indices are reported to assess and compare our platform with other algorithms including U-Net and cvi42, which is used in clinics. In addition, we trained and validated a deep fully convolutional network (FCN) on a dataset, consisting of \(64\) pediatric subjects with complex CHD, which we made publicly available. To mitigate this issue, we devised a novel method that uses a generative adversarial network (GAN) to synthetically augment the training dataset via generating synthetic CMR images and their corresponding chamber segmentations. Training artificial intelligence (AI) algorithms for CMR analysis requires large annotated datasets, which are not readily available for pediatric subjects and particularly in CHD patients. Thus, an automated and accurate segmentation platform exclusively dedicated to pediatric CMR images can significantly improve the clinical workflow, as the present work aims to establish. However, although CMR data facilitates reliable analysis of cardiac function and anatomy, clinical workflow mostly relies on manual analysis of CMR images, which is time consuming. Cardiovascular magnetic resonance (CMR) imaging offers non-invasive and non-ionizing assessment of CHD patients. For the growing patient population with congenital heart disease (CHD), improving clinical workflow, accuracy of diagnosis, and efficiency of analyses are considered unmet clinical needs.
